-Search query
-Search result
Showing all 23 items for (author: davies & km)

EMDB-24210: 
The 3D structure and in situ arrangements of CatSper channel from the cryo-electron tomography and subtomographic average of mouse wild type sperm flagella
Method: subtomogram averaging / : Zhao Y, Wang H, Wiesehoefer C, Shah NB, Reetz E, Hwang J, Huang X, Lishko PV, Davies KM, Wennemuth G, Nicastro D, Chung JJ

EMDB-26206: 
The 3D structure and in situ arrangements (Forward slash) of CatSper channel from the cryo-electron tomography and subtomographic average of mouse efcab9 mutant sperm flagella
Method: subtomogram averaging / : Zhao Y, Wang H, Wiesehoefer C, Shah NB, Reetz E, Hwang J, Huang X, Lishko PV, Davies KM, Wennemuth G, Nicastro D, Chung JJ

EMDB-26207: 
The 3D structure and in situ arrangements (Backslash) of CatSper channel from the cryo-electron tomography and subtomographic average of mouse efcab9 mutant sperm flagella
Method: subtomogram averaging / : Zhao Y, Wang H, Wiesehoefer C, Shah NB, Reetz E, Hwang J, Huang X, Lishko PV, Davies KM, Wennemuth G, Nicastro D, Chung JJ

EMDB-26483: 
Actin organization during clathrin-mediated endocytosis
Method: electron tomography / : Serwas D, Akamatsu M, Moayed A, Vegesna K, Vasan R, Hill JM, Schoeneberg J, Davies KM, Rangamani P, Drubin DG

EMDB-26484: 
In situ map of the clathrin hub
Method: subtomogram averaging / : Serwas D, Akamatsu M, Moayed A, Vegesna K, Vasan R, Hill JM, Schoeneberg J, Davies KM, Rangamani P, Drubin DG

EMDB-22829: 
Human Tom70 in complex with SARS CoV2 Orf9b
Method: single particle / : QCRG Structural Biology Consortium

EMDB-20207: 
Synthetic beta-carboxysome shell (T=4)
Method: single particle / : Sutter M, Laughlin TG, Sloan NB, Serwas D, Davies KM, Kerfeld CA

EMDB-20208: 
Synthetic beta-carboxysome shell (T=3)
Method: single particle / : Sutter M, Laughlin TG, Sloan NB, Serwas D, Davies KM, Kerfeld CA

EMDB-20209: 
Synthetic beta-carboxysome shell (prolate full)
Method: single particle / : Sutter M, Laughlin TG, Sloan NB, Serwas D, Davies KM, Kerfeld CA

EMDB-20210: 
Synthetic beta-carboxysome shell (T=4)
Method: single particle / : Sutter M, Laughlin TG, Sloan NB, Serwas D, Davies KM, Kerfeld CA

EMDB-0415: 
T.elongatus NDH (data-set 1)
Method: single particle / : Laughlin TG, Bayne A, Trempe JF, Savage DF, Davies KM

EMDB-0416: 
T.elongatus NDH Peripheral Arm Focus Map(data-set 1)
Method: single particle / : Laughlin TG, Davies KM

EMDB-0417: 
T.elongatus NDH Peripheral Arm Focus Map with NdhS(data-set 1)
Method: single particle / : Laughlin TG, Davies KM

EMDB-0418: 
T.elongatus NDH Peripheral Arm Focus Map without NdhS(data-set 1)
Method: single particle / : Laughlin TG, Davies KM

EMDB-0419: 
T.elongatus NDH Peripheral Arm Focus Map with X-cofactor(data-set 1)
Method: single particle / : Laughlin TG, Davies KM

EMDB-0420: 
T.elongatus NDH Peripheral Arm Focus Map without X-cofactor(data-set 1)
Method: single particle / : Laughlin TG, Davies KM

EMDB-0425: 
T.elongatus NDH (data-set 2)
Method: single particle / : Laughlin TG, Bayne A, Trempe JF, Savage DF, Davies KM

EMDB-3559: 
mitochondrial ATP synthase dimer from Euglena gracilis
Method: subtomogram averaging / : Muehleip AW, Davies KM, Kuehlbrandt W

EMDB-3560: 
mitochondrial ATP synthase dimer from Trypanosoma brucei
Method: subtomogram averaging / : Muehleip AW, Kuehlbrandt W, Davies KM

EMDB-3441: 
Subtomogram average of the mitochondrial ATP synthase dimer from the ciliate Paramecium tetraurelia
Method: electron tomography / : Muehleip AW, Kuehlbrandt W, Davies KM

EMDB-2982: 
Sub-tomogram average of a mammalian F-type ATP synthase monomer
Method: subtomogram averaging / : Jiko C, Davies KM, Shinzawa-Itoh K, Tani K, Maeda S, Mills DJ, Tsukihara T, Fujiyoshi Y, Kuehlbrandt W, Gerle C

EMDB-2852: 
Electron cryo-microscopy of mitochondrial ATP synthase dimers
Method: single particle / : Allegretti M, Klusch N, Mills DJ, Vonck J, Kuehlbrandt W, Davies KM
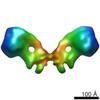
EMDB-2161: 
Structure of the yeast F1Fo-ATP synthase dimer
Method: subtomogram averaging / : Davies KM, Kuehlbrandt W
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
